Our Research
Adaptive Immune Receptor Discovery
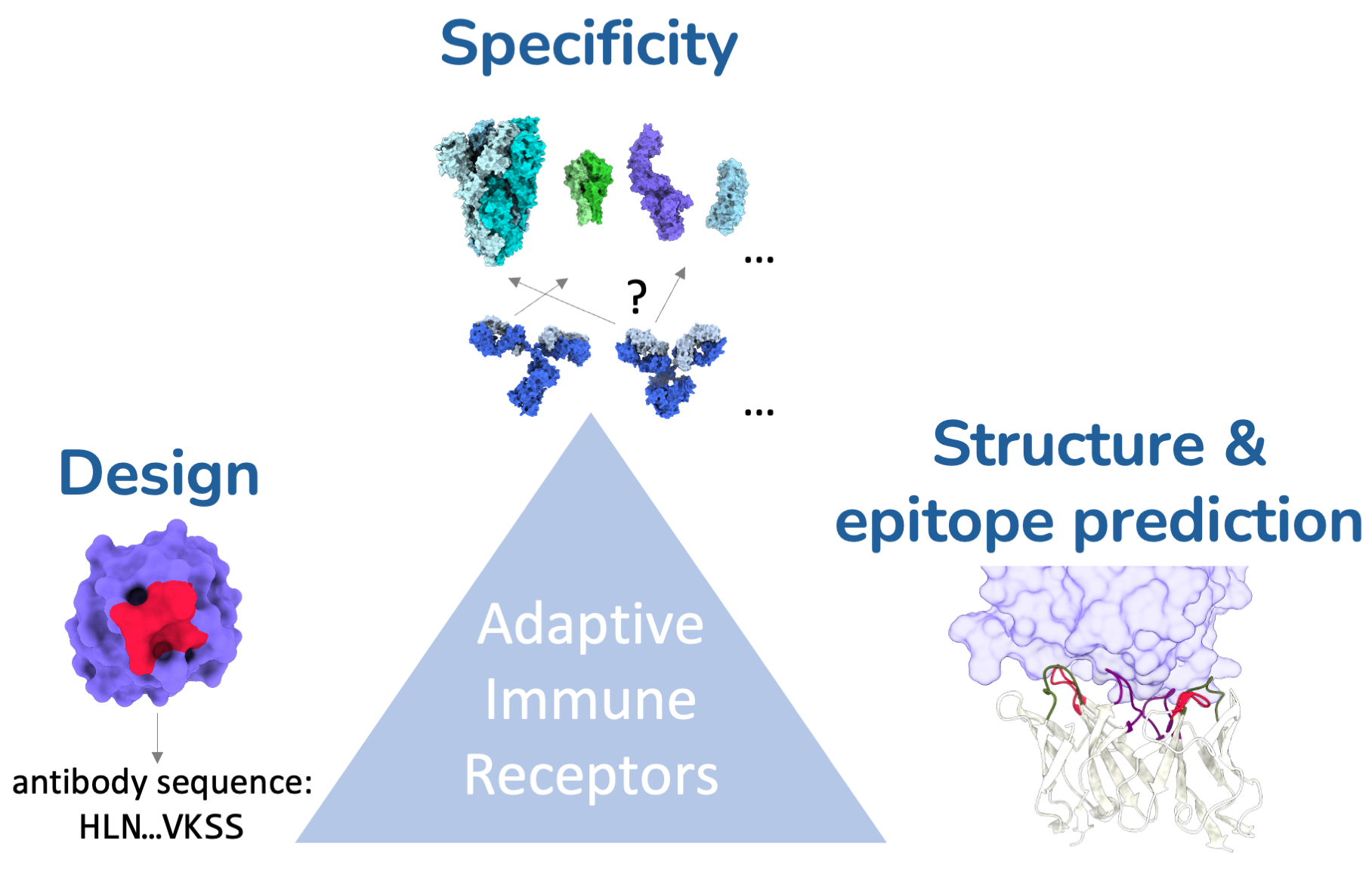
We develop deep learning methods for structure-based discovery of adaptive immune receptors, including antibodies, nanobodies, and T-cell receptors. Our goal is to turn large immune repertoire datasets into mechanistic insight about antigen recognition, epitope specificity, and immune protection.
- AIR-antigen structure modeling
- Specificity prediction
- Epitope-specific nanobody design

Integrative Approach
We develop integrative and hierarchical approaches for modeling large protein assemblies by combining deep learning-based structure prediction with experimental restraints and assembly algorithms.
- Pairwise interactions predicted by AlphaFold-based models are assembled into large multi-subunit complexes
- Combinatorial assembly enables modeling complexes with diverse stoichiometries, incomplete structural coverage, and multiple possible architectures
- Crosslinking mass spectrometry and other experimental restraints can be integrated to improve accuracy and confidence
- Our methods expand structural coverage for challenging assemblies that are difficult to resolve using end-to-end prediction alone
Variant Effect Prediction
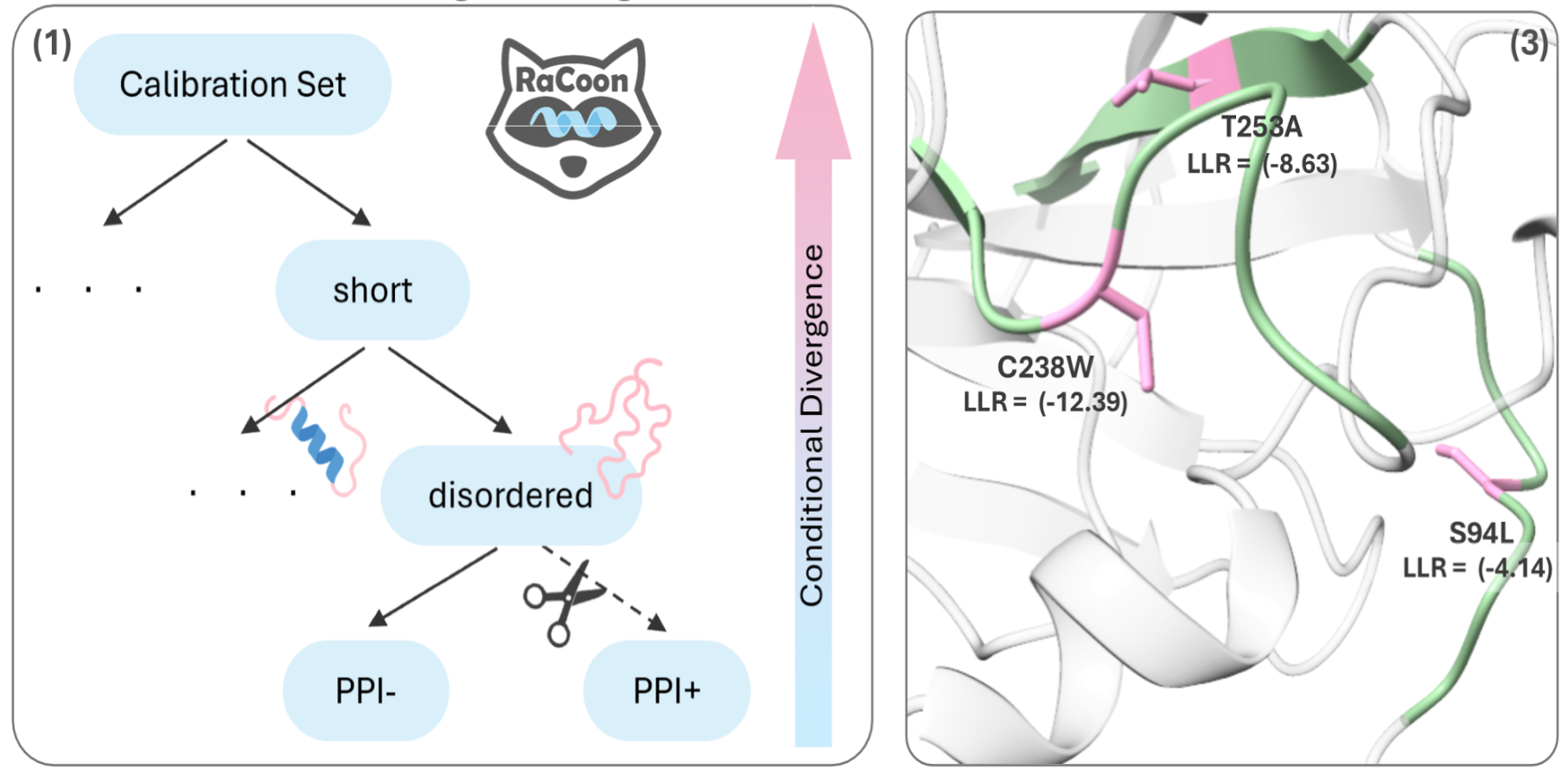
We develop machine learning approaches for interpreting missense variants at both the protein and residue levels, with an emphasis on accuracy, calibration, and biological interpretability. Our goal is to make variant effect prediction more reliable for gene discovery, rare disease studies, and clinical genomics.
- Protein-level prioritization of rare missense variants using deep learning scores and structural analysis
- Residue-aware calibration of pathogenicity predictions to improve reliability across distinct variant subgroups
- Interpretation of variant effects in biological pathways and disease-associated proteins
Rhodopsin Engineering
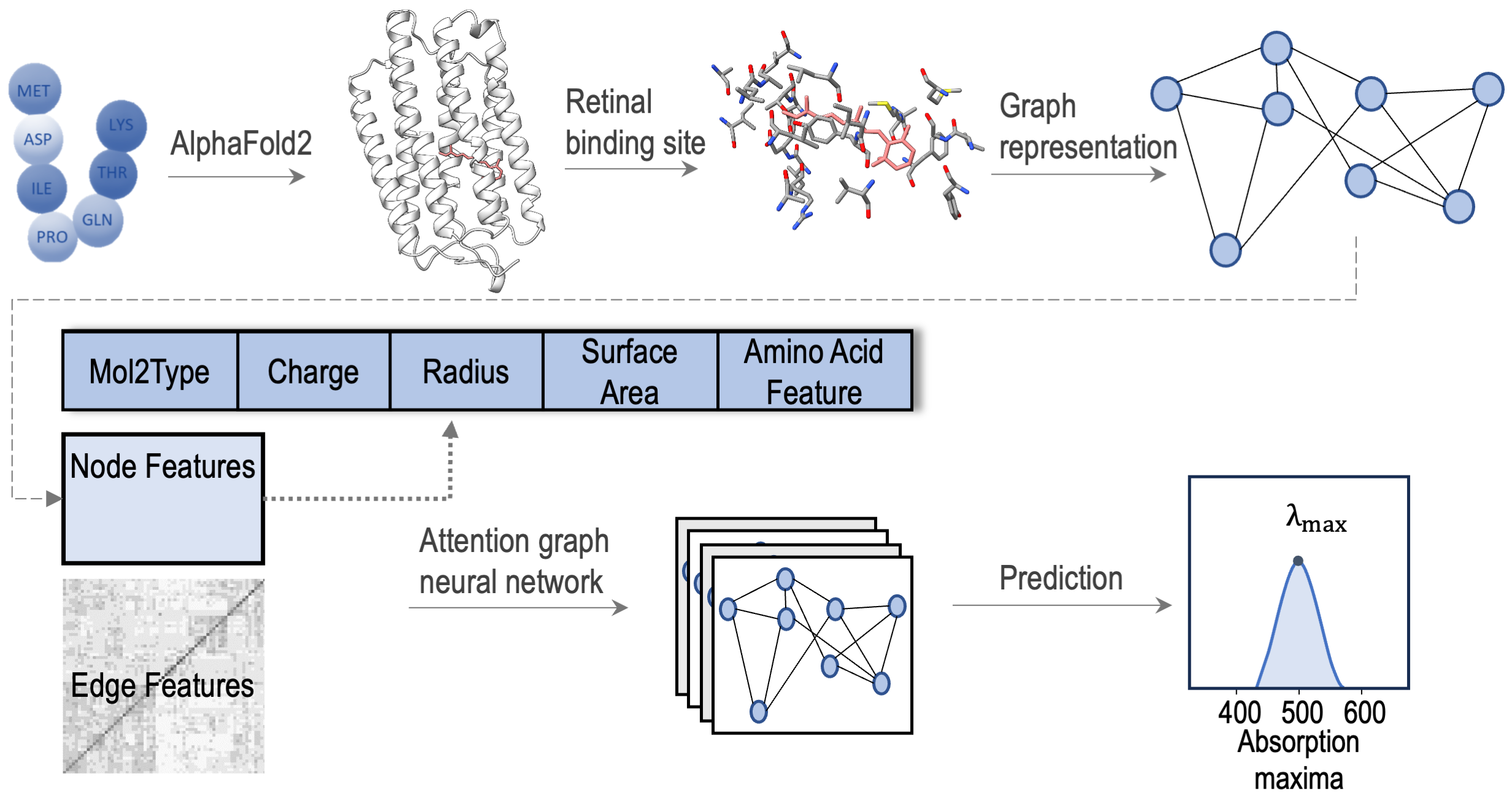
We develop AI methods for predicting and engineering functional properties of microbial rhodopsins and channelrhodopsins. Building on structure prediction and geometric deep learning, we study how sequence and three-dimensional environment shape spectral tuning and channel properties in light-activated membrane proteins.
- Prediction of rhodopsin absorption maxima from sequence and structure
- Integration of geometric deep learning with protein language models for function prediction
- Computational design of red-shifted and functionally optimized rhodopsins
Dynamics Modeling
Dynamics modeling is required to understand structural states, their populations, and the transition pathways between them.
- Sampling relevant states
- Multi-state model enumeration
- Scoring against experimental data
- Determining population weights
