פרופסור, ראש החוג לביולוגיה חישובית
בית הספר להנדסה ולמדעי המחשב, בית הספר לרפואה
האוניברסיטה העברית בירושלים
 |
פרופסור, ראש החוג לביולוגיה חישוביתבית הספר להנדסה ולמדעי המחשב, בית הספר לרפואההאוניברסיטה העברית בירושלים
Professor, Head of the Computational Biology ProgramSchool of Computer Science and Engineering, Faculty of MedicineThe Hebrew University of Jerusalem, Israel |
Computational BiologyDNA Methylation of cell-free DNA, Computational Epigenomics, Enhancers and Gene Regulation, Machine Learning |
Prof. Tommy Kaplan
School of Computer Science and Engineering
The Hebrew University
Jerusalem 9190401
Israel
E-mail: tommy.kaplan@mail.huji.ac.il
Office: Rothberg B-529
Phone/Fax: +972-2-5494506
| Netanel Loyfer*, Judith Magenheim*, Ayelet Peretz*,
Gordon Cann, Joerg Bredno, Agnes Klochendler, Ilana Fox-Fisher,
Sapir Shabi-Porat, Merav Hecht, Tsuria Pelet, Joshua
Moss, Zeina Drawshy, Hamed Amini, Patriss Moradi, Sudharani
Nagaraju, Dvora Bauman, David Shveiky, Shay Porat, Gurion Rivkin,
Omer Or, Nir Hirshoren, Einat Carmon, Alon Pikarsky, Abed
Khalaileh, Gideon Zamir, Ronit Grinbaum, Machmud Abu Gazala, Ido
Mizrahi, Noam Shussman, Amit Korach, Ori Wald, Uzi Izhar, Eldad
Erez, Vladimir Yutkin, Yaacov Samet, Devorah Rotnemer Golinkin,
Kirsty L. Spalding, Henrik Druid, Peter Arner, A.M. James Shapiro,
Markus Grompe, Alex Aravanis, Oliver Venn, Arash Jamshidi, Ruth
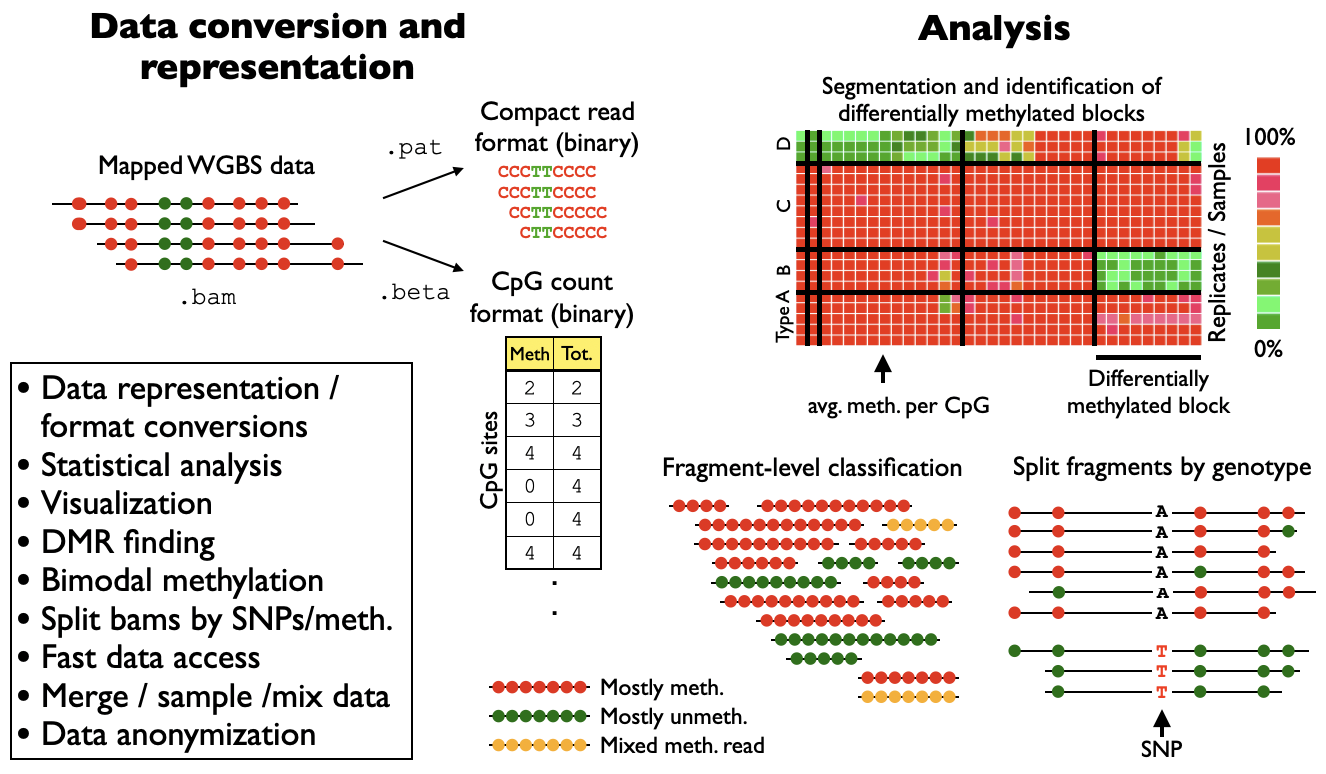
Shemer, Yuval Dor*, Benjamin Glaser*, and Tommy Kaplan* A DNA methylation atlas of normal human cell types Nature, 2023 [GEO GSE186458 | poster | הארץ | הידען | Research Highlight] Genome browser sessions: hg19 | hg38 Software: wgbstools | UXM fragment-level deconvolution |
| Miri Varshavsky, Gil Harari, Benjamin Glaser, Yuval Dor, Ruth Shemer, and Tommy Kaplan Accurate age prediction from blood using a small set of DNA methylation sites and a cohort-based machine learning algorithm Cell Reports Methods, 2023 [GP-age epigenetic clock | poster | GEO GSE207605] |
| Megan E. McNamara*, Netanel Loyfer*, Amber J. Kiliti, Marcel O. Schmidt,
Sapir Shabi-Porat, Sidharth Jain, Sarah Martinez Roth, A. Patrick McDeed IV, Nesreen Shahrour,
Elizabeth Ballew, Yun-Tien Lin, Heng-Hong Li, Anne Deslattes Mays, Sonali Rudra, Anna T. Riegel,
Keith Unger*, Tommy Kaplan*, and Anton Wellstein*
Circulating cell-free methylated DNA reveals tissue-specific, cellular damage from radiation treatment
JCI Insight, 2023 Software: wgbstools | fragment-level Markov-model deconvolution |
| Ayelet Peretz*, Netanel Loyfer*,
Sheina Piyanzin, Bracha Lea Ochana, Daniel Neiman, Judith Magenheim, Agnes Klochendler, Zeina Drawshy,
Ilana Fox-Fisher, Ori Fridlich, Joshua Moss, Daniel Cohen, Hai Zemmour, Gordon Cann, Joerg Bredno,
Oliver Venn, Batia Avni, Tural Alekberli, Yaacov Samet, Amit Korach, Ori Wald, Vladimir Yutkin, Uzi Izhar,
Nir Pillar, Markus Grompe, Zvi Fridlender, Ariel Rokach, David Planer, Giora Landesberg,
Benjamin Glaser, Ruth Shemer, Tommy Kaplan and Yuval Dor The DNA methylome of human vascular endothelium and its use in liquid biopsies Med, 2023 |
| Jonathan Rosenski, Sagiv Shifman, and Tommy Kaplan Predicting gene knockout effects from expression data BMC Medical Genomics, 2023 [github] |
 |
Aleks Danov, Ofir Segev, Avi Bograd, Yedidya Ben Eliyahu, Noam Dotan, Tommy Kaplan*, and Asaf Levy* Toxinome - The Bacterial Protein Toxin Database mBio, 2023 [Toxinome: the Bacterial toxin database] |
| Mohammad Jaber*, Ahmed Radwan*, Netanel Loyfer*, Mufeed
Abdeen, Shulamit Sebban, Thorsten Kolb, Marc Zapatka, Kirill
Makedonski, Aurelie Ernst, Tommy Kaplan*, and Yossi
Buganim* Comparative Parallel Multi-Omics Analysis During the Induction of Pluripotent and Trophectoderm States Nature Communications, 2022 [GEO GSE171127 | הארץ | הידען ] |
| Asael Lubotzky, Hai Zemmour, Daniel Neiman, Marc Gotkine,
Netanel Loyfer, Sheina Piyanzin, Bracha-Lea Ochana, Roni
Lehmann-Werman, Daniel Cohen, Joshua Moss, Judith Magenheim,
Maureen F. Loftus, Lauren Brais, Kimmie Ng, Raul Mostoslavsky,
Brian M. Wolpin, Aviad Zick, Myriam Maoz, Albert Grinshpun, Anatoli
Kustanovich, Chen Makranz, Jonathan E. Cohen, Tamar Peretz, Ayala
Hubert, Mark Temper, Azzam Salah, Shani Avniel-Polak, Simona
Grozinsky-Glasberg, Kirsty L. Spalding, Ariel Rokach, Tommy
Kaplan, Benjamin Glaser, Ruth Shemer, and Yuval Dor Liquid biopsy reveals collateral tissue damage in cancer JCI Insight, 2022 |
| Ilana Fox-Fisher, Sheina Piyanzin, Bracha Lea Ochana, Agnes
Klochendler, Judith Magenheim, Ayelet Peretz, Netanel
Loyfer, Joshua Moss, Daniel Cohen, Yaron Drori, Nehemya
Friedman, Michal Mandelboim, Marc E Rothenberg, Julie M Caldwell,
Mark Rochman, Arash Jamshidi, Gordon Cann, David Lavi, Tommy
Kaplan, Benjamin Glaser, Ruth Shemer, and Yuval Dor Remote immune processes revealed by immune-derived circulating cell-free DNA eLife, 2021 |
 |
Megan E. Barefoot, Netanel Loyfer, Amber J. Kiliti, A.
Patrick McDeed IV, Tommy Kaplan, and Anton Wellstein Detection of Cell Types Contributing to Cancer From Circulating, Cell-Free Methylated DNA Frontiers in Genetics, 2021 |
| Naomi Habib, Cristin McCabe, Sedi Medina, Miriam Varshavsky,
Daniel Kitsberg, Raz Dvir-Szternfeld, Gilad Green, Danielle Dionne,
Lan Nguyen, Jamie L. Marshall, Fei Chen, Feng Zhang, Tommy
Kaplan, Aviv Regev, and Michal Schwartz Disease-associated astrocytes in Alzheimer’s disease and aging Nature Neuroscience, 2020 [GEO GSE143758] |
| Yifat Brill-Karniely, Dvir Dror, Tal Duanis-Assaf, Yoel
Goldstein, Ouri Schwob, Talya Millo, Natalie Orehov, Tal Stern,
Mohammad Jaber, Netanel Loyfer, Margarita Vosk-Artzi, Hadar
Benyamini, Diane Bielenberg, Tommy Kaplan, Yosef Buganim,
Meital Reches and Ofra Benny Triangular correlation (TrC) between cancer aggressiveness, cell uptake capability, and cell deformability Science Advances, 6(3):eaax2861, 10.1126/sciadv.aax2861 [GEO GSE137316, GSE135193] |
| Ronen Sadeh, Israa Sharkia, ... , Tommy Kaplan, Ruth
Shemer, David Planer, Eithan Galun, Benjamin Glaser, Aviad Zik,
Yuval Dor, and Nir Friedman ChIP-seq of plasma cell-free nucleosomes identifies gene expression programs of the cells of origin Nature Biotechnology, 2021 [הארץ] |
| Hana Benchetrit, Mohammad Jaber, Valery Zayat, Shulamit Sebban,
Avital Pushett, Kirill Makedonski, Zvi Zakheim, Ahmed Radwan, Noam
Maoz, Rachel Lasry, Noa Renous, Michal Inbar, Oren Ram, Tommy
Kaplan, and Yosef Buganim Direct Induction of the Three Pre-implantation Blastocyst Cell Types from Fibroblasts Cell Stem Cell, 2019, 10.1016/j.stem.2019.03.018 [GEO ] |
| Joshua Moss, Judith Magenheim, Daniel Neiman, Hai
Zemmour, Netanel Loyfer, Amit Korach, Yaacov Samet, Myriam
Maoz, Henrik Druid, Peter Arner, Keng-Yeh Fu, Endre Kiss, Kirsty L.
Spalding, Giora Landesberg, Aviad Zick, Albert Grinshpun, AM James
Shapiro, Markus Grompe, Avigail Dreazan Wittenberg, Benjamin
Glaser, Ruth Shemer*, Tommy Kaplan*, and Yuval Dor* Comprehensive human cell-type methylation atlas reveals origins of circulating cell-free DNA in health and disease Nature Communications, 2018, 9:5068 [GEO | github meth atlas] |
| Dikla Cohn, Or Zuk*, and Tommy Kaplan* Enhancer Identification using Transfer and Adversarial Deep Learning of DNA Sequences bioRxiv, 2018 [enhancer_CNN github | enhancer datasets] |
| Gil Ron, Yuval Globerson, Dror Moran, and
Tommy Kaplan Promoter-Enhancer Interactions Identified from Hi-C Data using Probabilistic Models and Hierarchical Topological Domains Nature Communications, 2017, 8(1):2237 [PSYCHIC github | poster] |
| Yuval Malka, Avital Steiman-Shimony, Eran Rosenthal,
Leonor Cohen-Daniel, Eliran Arbib, Liron Argaman, Hanah Margalit,
Tommy Kaplan* and Michael Berger* Post-transcriptional 3'-UTR cleavage of mRNA transcripts generates thousands of stable uncapped autonomous RNA fragments Nature Communications, 2017, 8(1):2029 [GEO ] |
| Michael Klutstein*, Joshua Moss*, Tommy Kaplan,
and Howard Cedar Contribution of epigenetic mechanisms to variation in cancer risk among tissues PNAS, 2017 |
| Xiao-Yong Li, Melissa M. Harrison, Jacqueline E. Villalta,
Tommy Kaplan*, and Michael B. Eisen* Establishment of regions of genomic activity during the Drosophila maternal to zygotic transition eLife, 2014;3:e03737 [Digest | GEO | Genome Browser] |
| Axel Visel,... Dalit May,... Tommy Kaplan,... Edward M.
Rubin, Ivan Ovcharenko, Len A. Pennacchio, John L. R.
Rubenstein A High-Resolution Enhancer Atlas of the Developing Telencephalon Cell, 2013, 10.1016/j.cell.2012.12.041 |
 |
Tommy Kaplan and Nir Friedman Gene Expression: Running to Stand Still Nature, 2012, 484:171-172 [News and Views item of Lickwar et al's Genome-wide protein-DNA binding dynamics suggest a molecular clutch for transcription factor function] |
 |
Dalit May, Matthew J. Blow, Tommy Kaplan, David J.
McCulley, Brian C. Jensen, Jennifer A. Akiyama, Amy Holt, Ingrid
Plajzer-Frick, Malak Shoukry, Crystal Wright, Veena Afzal, Paul C.
Simpson, Edward M. Rubin, Brian L. Black, James Bristow, Len A.
Pennacchio, and Axel Visel Large-Scale Discovery of Enhancers from Human Heart Tissue Nature Genetics, 2012, 44:89-93 [ Abstract | Supp. Information ] |
 |
Melissa M. Harrison*, Xiao-Yong Li*, Tommy Kaplan*,
Michael R. Botchan and Michael B. Eisen Zelda binding in the early Drosophila melanogaster embryo marks regions subsequently activated at the maternal-to-zygotic transition PLoS Genetics, 2011, 7(10): e1002266 [ Why Zelda is cool? | HHMI news | Grizzly Peak fitting | Nat. Rev. Gen In Brief | F1000 ] |
 |
Tommy Kaplan, Xiao-Yong Li, Peter Sabo, John A.
Stamatoyannopoulos, Mark D. Biggin, and Michael B. Eisen Quantitative Models of the Mechanisms that Control Genome-Wide Patterns of Transcription Factor Binding During Early Drosophila Development PLoS Genetics, 2011, 7(2): e1001290 [Book chapter] |
| Moran Yassour*, Tommy Kaplan*, Hunter B. Fraser, Joshua
Z. Levine, Jenna Pfiffner, Xian Adiconis, Gary Schroth, Shujun Luo,
Irina Khrebtukova, Andreas Gnirke, Chad Nusbaum, Dawn-Anne
Thompson, Nir Friedman, and Aviv Regev Ab initio Construction of a Eukaryotic Transcriptome by Massively Parallel mRNA Sequencing [Supp. Data] PNAS, 2009, 106(9):3264-9 |
 |
Andrew P. Capaldi, Tommy Kaplan, Ying Liu, Naomi Habib,
Aviv Regev, Nir Friedman and Erin K. O'Shea Structure and Function of a Transcriptional Network Activated by the MAPK Hog1 Nature Genetics, 2008, 40:1300-6 [Abstract | Supp. information | Supp. webpage | ChIP-chip Peak Fitting Algorithm | F1000 | NBT's news and views ] |
 |
Micheal F Dion*, Tommy Kaplan*, Minkyu Kim, Stephen
Buratowski, Nir Friedman, and Oliver J. Rando Dynamics of Replication-Independent Histone Turnover in Budding Yeast Science, 2007, 315(5817):1405-8 [ Abstract | Poster | F1000 ] |
 |
Chih Long Liu*, Tommy Kaplan*, Minkyu Kim, Stephen
Buratowski, Stuart L. Schreiber, Nir Friedman, and Oliver J.
Rando Single-Nucleosome Mapping of Histone Modifications in S. cerevisiae PLoS Biology, 2005, 3(10):e328 [ Interactive Map | F1000 ] |
 |
Tommy Kaplan, Nir Friedman and Hanah Margalit ab initio Prediction of Transcription Factor Targets using Structural Knowledge PLoS Comp. Biol., 2005, 1(1):e1 |
| Yoseph Barash, Gal Elidan, Nir Friedman, and Tommy
Kaplan Modeling Dependencies in Protein-DNA Binding Sites Proc 7th Ann Int Conf in Comp Mol Bio (RECOMB), 2003 |
 |
Netanel
Loyfer Computational Analysis of whole-genome bisulfite-seq DNA methylation data (Best Poster Awards in Hebrew University's Open Day 2019, Broad-ISF conference 2019) [PhD] |
 |
Jonathan
Rosenski Machine learning analysis of parental imprinting in DNA methylation data [PhD] |
 |
Roy
Mizrachi DNA methylation in forensics sciences [MSc] |
 |
Mika
Schechter Predicting 3D genome folding using computational methods [MSc] |
 |
Roni Schwartz DNA methylation hotspots and nucleosomes [MSc] |
 |
Daniel Nudelman Deep learning predictions of age from DNA methylation [MSc] |
 |
Miri
Varshavsky GP-Age: Accurate age prediction from blood using few methylation sites [Poster] (Best Poster Award in Ilanit 2023) [MSc] |
 |
Hadar
Mulian Semi-constrained non-negative matrix factorization models for cell-free DNA data [MSc, CS/LS program] |
 |
Sapir
Shabi Modeling dependencies in cell-free DNA methylation data (Best Poster Award at the Broad-ISF conference 2019) [MSc, CS/LS program] |
 |
Noam
Glausiusz Modeling dependencies in DNA methylation sequencing data [MSc, CS program, with Prof. Gal Elidan] |
 |
Ido
Lokay Phylogenetic Mixture Model for DNA Methylation Patterns Formation During Cell Differentiation [MSc, CSE program] |
 |
Josh
Moss Computational Aspects of Cell-Free DNA (Kaye Innovation Award, 2019) [MD/PhD program, with Prof. Yuval Dor] |
 |
Lior
Ziv Computational analysis of short- and long-read genomics in the brain [MSc, CS/LS program, with Dr. Ami Citri] |
 |
Guy Kelman,
PhD Comparative analysis in Bat and Mouse identifies wing-specific genes and regulatory regions (Best Poster Awards in IBS 2017 and Hebrew University's Open Day 2017) [Postdoc] Guy is our local Batman, using mRNA-seq and ChIP-seq data to indentify Enhancers in developing Bat wings. |
 |
David
Taub Classification of Enhancer sequencing in multiple tissues using k-order Markovian models [MSc, CS program] David is developing structured Hidden Markov Models (sHMMs) of DNA sequences, using high-order probabilistic emission models. |
 |
Yuval
Globerson Analyzing 3D genomic data using Spectral methods and other Computational models. [MSc, CS program] |
 |
Paz
Bunis Deep Learning models of DNA sequences for multi-label multi-class Enhancer classification [MSc, CS program] Convolutional Neural Networks for Enhancer classification based on DNA shape and seqeunce. |
 |
Dikla
Cohn Enhancer Identification using Transfer and Adversarial Deep Learning of DNA Sequences [github enhancer_CNN] [MSc, CS] |
 |
Gil Ron Finding Hierarchies in Topological Domains using a Unified Bayesian Interaction Model [MSc, CS] |
 |
Eran
Rosenthal Computational models of large-scale genomics data [MSc, CS/LS program] |
 |
Shira
(Strauss) Ginzburg, MSc Genome-wide 3D maps of regulatory interactions in the mouse developing forebrain [MSc, CS/LS program] |
 |
Inbar Naor,
MSc Classification of Putative Regulatory Enhancers from DNA sequences [MSc, CS] Machine Learning sequence classifiers to identify regulatory DNA regions and decypher their function. |